Dhuha Oday Hassan 1 , Alaa Hassan Jawad 2 , Shakir H. Mohammed Al. Alwany 3
Authors affiliations:
Background & objective: Patients with chronic myeloid leukemia (CML) remain at a risk of hepatitis B virus (HBV) reactivation during tyrosine kinase inhibitor (TKI) therapy. Programmed death-ligand 1 (PD-L1) polymorphisms may influence immune evasion and disease pathogenesis; however, their interaction with HBV in CML, particularly in Iraqi populations, remains uncharted. We conducted this study to investigate PD-L1 genetic variations, serum PD-L1 levels, and HBV virus infection.
Methodology: This was a case-control study of 120 patients with CML and 100 healthy controls (AHC). Serum PD-L1 and HBV markers were quantified by ELISA Genomic DNA was extracted, and specific regions of the PD-L1 gene and HBV DNA were amplified by PCR. PD-L1 amplicons were sequenced to identify genetic variants.
Results: CML patients exhibited significantly lower serum PD-L1 levels than controls (mean 268.8 ± 89 vs. 470.6 ± 288, P = 0.0032). No HBV DNA was detected in any participant. A novel C>T substitution in PD-L1 was identified, with TT and CT genotypes being more frequent in CML patients than in controls. The T allele showed a borderline significant association with a reduced CML risk (P = 0.01, OR = 0.22, 95% CI: 0.06–0.73).
Conclusion: The PD-L1 T allele may confer protection against CML, while suppression of serum PD-L1 suggests distinct regulatory mechanisms. The absence of HBV highlights regional epidemiological patterns. PD-L1 genotyping can inform risk stratification of patients with Iraqi CML.
Abbreviations: CML: chronic myeloid leukemia, HBV: hepatitis B virus, PD-L1: TKI: tyrosine kinase inhibitor
Keywords: Chronic myeloid leukemia, Programmed Death-Ligand 1, Hepatitis B virus, Immune checkpoint.
Citation: Hassan DO, Jawad AH, Al-Alwany SHM. Immunogenetic study of programmed death-ligand 1 variation and hepatitis B virus in Iraqi patients with chronic myeloid leukemia. Anaesth. pain intensive care 2025;29(7):749-755. DOI: 10.35975/apic.v29i7.2967
Received: May 09, 2025; Revised: October 26, 2025; Accepted: January 01, 2025
Chronic myeloid leukemia (CML) is a hematopoietic stem cell (HSC) disorder, like all leukemias, and is a type of cancer that starts in the blood-forming cells of the bone marrow and invades the blood. It is also known as chronic myelogenous leukemia. It is a myeloproliferative neoplasm with an incidence of 1–2 cases per 100,000 adults.1 It is defined by the presence of the Philadelphia chromosome (Ph), which results from reciprocal translocation between chromosomes 9 and 22 [t (9;22] in a single bone marrow cell during cell division. Part of chromosome 9 attaches to chromosome 22, and part of chromosome 22 attaches to chromosome 9. Chromosome 9 was longer than normal, and chromosome 22 was shorter than normal. Abnormal chromosome 22 is known as the “Philadelphia chromosome, which gives rise to a BCR-ABL1 fusion gene.2
CML is classified into three distinct clinical phases: the chronic, accelerated, and blast phases, which are defined by the proportion of immature myeloid blasts in the blood and bone marrow, along with secondary cytogenetic or clinical features.3,4 In the chronic phase, patients typically exhibit <10% blasts and retain partial myeloid differentiation, often presenting with leukocytosis, thrombocytosis, and mild anemia.5,6 Progression to the accelerated phase is marked by 10–19% blasts, ≥20% peripheral basophils, or clonal cytogenetic evolution, reflecting increased genomic instability.7 The blast phase, resembling acute leukemia, is characterized by ≥20% blasts, extramedullary proliferation, or lymphoid blast crisis, indicating a complete loss of differentiation capacity.8
Programmed death ligand 1 (PD-L1), also called (CD274) located on chromosome 9p24.1 and is expressed in non-hematopoietic cells, hematopoietic cells, and antigen-presenting cells (APC).9 An important role of PD-L1 is in various malignancies, where it can attenuate the host immune response to tumor cells.10 PD-L1 expressed by cancer cells binds to PD-1 on the surface of T cells, thereby inhibiting T cell activation and leading to cancer immunity, which allows the tumor to grow without being attacked by the immune system.11
Hepatitis B virus (HBV) infection lasts 45-180 days, most commonly 60-90 days.12 It contains a circular, relaxed DNA (rcDNA) of approximately 3.2 kb in length with a complete minus strand and an incomplete plus strand, which represents the smallest DNA viruses infecting humans. Infected blood or other bodily fluids are involved in the entry of the virus into the body. Contaminated needles or medical equipment, contaminated blood transfusions, unprotected intercourse with an infected individual, or vertical transmission from mother to child during pregnancy or delivery are all possible routes of transmission.13 According to several studies, cytotoxic chemotherapy and immunosuppressive drugs have the potential to hasten the reactivation of chronic HBV infection, putting patients undergoing these treatments at an increased risk for adverse effects.14
Despite established evidence that patients with CML treated with tyrosine kinase inhibitors (TKIs) are at risk of HBV reactivation, standardized HBV screening or monitoring protocols are lacking. Furthermore, while PD-L1 is a critical immune checkpoint implicated in cancer immune evasion, the role of specific PD-L1 gene polymorphism and their potential interaction with HBV infection in modulating CML susceptibility or pathogenesis remains poorly characterized, particularly within the Iraqi population.15,16
This study aimed to address these gaps by investigating PD-L1 genetic variations, PD-L1 serum levels, and HBV infection status as potential risk or protective factors in Iraqi patients with CML
This study employed a cross-sectional case-control design. A total of 220 serum specimens were collected, comprising 120 patients diagnosed with CML, and 100 apparently healthy control (AHC) individuals. CML patients’ specimens were obtained from the Iraqi Hematology Center, whereas AHC specimens were collected from general hospitals within the Middle Euphrates region of Iraq. CML patients (aged 20–60 years) met the following criteria: (1) BCR-ABL1+ chronic-phase diagnosis, (2) no active infections at enrollment, (3) no prior allogeneic transplantation, and (4) absence of HIV/HBV/HCV co-infection or concurrent malignancies. AHCs were screened via questionnaire, physical examination, complete blood count (CBC), and C-reactive protein (CRP) testing to exclude immunocompromised states, chronic diseases, or recent infections. All specimens were collected between October 2024 and January 2025.
For serum preparation, approximately 3 mL of venous blood was drawn from each participant into gel-containing serum separators. Samples were centrifuged according to standard protocols to separate the serum, which was then immediately aliquoted and stored at -20°C until subsequent analysis. The serum levels of HBV markers and PD-L1) were quantitatively assessed for both the CML group and the AHC group, using commercially available Enzyme-Linked Immunosorbent Assay (ELISA) kits (BT LAB, China), following the manufacturer's instructions.
2.1. DNA Extraction
Total genomic DNA was isolated from approximately 5 mg of whole blood collected from CML and AHC patients. Samples were immediately transferred to sterile tubes, refrigerated, and homogenized using a hand-held homogenizer in 1.5 mL Eppendorf tubes. Extraction was performed using the G-Spin Total DNA Extraction Kit (iNtRON Biotechnology Co., Korea), and the purified DNA was stored at −20°C. Viral DNA was isolated using a Viral DNA Extraction Kit (iNtRON Biotechnology Co., Korea) under identical storage conditions.
2.2. Primer Design and PCR Amplification
Gene-specific primers were designed for PD-L1 (forward: 5′-TTCCCACTCCCCACTGTTAG-3′; reverse: 5′-GCCCACAGCCACATAAACTT-3′) and HBV (forward: 5′-ACATGGAGAACATCGCATCA-3′; reverse: 5′-AGGACAAACGGGCAACATAC-3′). PCR reactions were conducted in a 25 μL volume comprising 12.5 μL master mix, 5 μL DNA template, 1 μL of each primer, and 5.5 μL nuclease-free water. Amplification was performed using a conventional thermal cycler (Germany) under uniform conditions for both targets: initial denaturation at 95°C for 5 min; 35 cycles of denaturation (95°C for 1 min), annealing (58°C for 45 s), and extension (72°C for 2 min), followed by a final extension at 72°C for 5 min.
Amplicons were resolved using 1.5% agarose gel electrophoresis and visualized under UV transillumination. The PD-L1 PCR products were sequenced using the forward primer via automated Sanger sequencing (Macrogen Inc., Seoul, South Korea). Sequence alignment and analysis were performed using Geneious Bioinformatics software (v2.0) against NCBI reference databases.
2.3. Statistical analysis
To assess the significance of the variables examined in this study, the chi-square test was used. All statistical analyses were conducted using SPSS program version 23. Statistical significance was set at P < 0.05. Hardy-Weinberg equilibrium was used to analyze PD-L1 gene polymorphism.
Table 1 compares the demographic and clinical parameters of the 120 CML patients (including 20 newly diagnosed and 100 treated cases) and 100 AHC. Age and sex distributions showed no significant differences (P = 0.06 for age; P = 0.059 for sex), with mean ages of 44 ± 13.3 years (new CML), 48 ± 11.43 years (treated CML), and 46 ± 12.9 years (AHC). However, newly diagnosed CML patients exhibited markedly elevated white blood cell (WBC) counts (173.5 ± 88.13 ×10⁹/L) compared to AHC (6.6 ± 0.75 ×10⁹/L; p<0.001). Platelet counts were also significantly higher in new CML patients (283 ± 138 ×10⁹/L) versus AHC (246.6 ± 56.3 ×10⁹/L; P < 0.05). Hemoglobin (Hb), packed cell volume (PCV), and BCR-ABL ratios did not differ significantly between the CML and AHC groups.
Table 2 shows significantly lower serum PD-L1 levels in CML patients (n = 120) than in apparently healthy controls (AHC; n = 100). The mean CTLA-4 concentration was 268.8 ± 89 in CML patients versus 470.6 ± 288 in the AHC group (P = 0.0032).
HBV-DNA was absent in all 120 CML patients tested via PCR. All samples were negative (100%), with no positive cases (0%). The P = 0.03 was inconsistent with the non-significant notation (Sign > 0.05), though the data confirm no HBV infection in the cohort. The AHC groups similarly tested negative, supporting the conclusion that HBV was undetectable in both study populations (Figure 1).

Figure 1: non- PCR detection of the HBV gene; in CML patients. M: A 100–1100 bp DNA ladder. After migrating into 2% agarose at 75V and 20 mA for 120 minutes, the PCR-amplified products were stained with ethidium bromide and placed in 15 µl per well
In the current study, the PD-L1 genetic sequences located on chromosome 9 were targeted. The 439 bp amplicon analyzed in this study represents part of the exon region of the PD-L1 molecule (Figure 2).
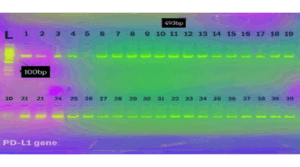
Figure 2: PCR detection of the PDL-1 gene; in CML patients. M: A 100–1100 bp DNA ladder. After migrating into 2% agarose at 75V and 20 mA for 120 minutes, the PCR-amplified products were stained with ethidium bromide and placed in 15 µl per well.
The NCBI BLASTn engine exhibited approximately 99.9% sequence similarity between the intended target reference sequences and the sample gene sequences. The alignment results of the 493 bp samples were compared to the reference DNA sequences (Figure 3). The mutation was observed at a specific position showing a C > T substitution.
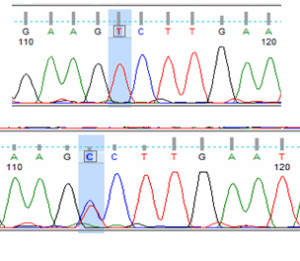
Figure 3: A Novel mutation in PD-L1 gene.
As illustrated in Figure 3, the polymorphism appeared in two distinct zygosity patterns among the sequenced samples: a homozygous (T/T) form and a heterozygous (C/T) form, which was confirmed through sequencing chromatograms. The chromatographic profiles demonstrated a clear peak separation, highlighting the single-nucleotide variation between the samples.
The present results showed that the DNA polymorphism distributions were as follows: CC, CT, and TT were 25%, 30%, and 45%, respectively, in patients with CML and 70%, 10%, and 20%, respectively, in the control group. There were no statistically significant differences (p< 0.05) between the different groups according to PD-L1 genotyping. Statistical analysis of CT indicated a borderline non-significant association (P = 0.08) with an odds ratio (OR) of 0.11 (95% confidence interval [CI]: 0.01–1.32), as shown in Table 4. Statistical analysis of the TT genotype indicated a borderline non-significant association (P = 0.05) with an OR of 0.15 (95% CI: 0.02–1.07), suggesting a potential but not statistically confirmed protective effect.
The C allele was present in 16 patients (40%) and 15 control alleles (75%) and served as the reference allele. The T allele was observed in 24 patients (60%) and 5 control alleles (25%). Statistical analysis showed a borderline significant association between the T allele and decreased disease risk (P = 0.01), with an OR of 0.22 (95% CI: 0.06–0.73) as shown in Table 3.
A novel partial sequence of the PD-L1 gene was identified, covering 439 bp and located on chromosome 9. The samples were collected from blood-derived leukocytes and submitted to the DDBJ/EMBL/GenBank databases under accession numbers LC867662, LC867663, and LC867664.
In this study, the genetic sequence of PD-L1 was analyzed. For the currently investigated 439 bp gene amplicon, BLASTn analysis revealed a high similarity with the reference human PD-L1 gene sequences available in NCBI, including coding and adjacent regulatory regions, according to the human genome annotation (GenBank version: LC867662.1: LC867663.1, and LC867664.1).
CML is the most common leukemia, in which the BCR-ABL1 fusion gene, which increases tyrosine kinase activity and inhibits apoptotic mechanisms, is responsible for the unchecked clonal proliferation of hematopoietic stem cells that characterizes CML, a myeloproliferative neoplasm. The hallmark of disease pathogenesis is the increase of immature granulocytes in the bone marrow and peripheral blood as a result of this.17–19
In the current study, the average age of the patients with a new diagnosis was 44 years, and the group had an average age of 51 years. However, the P-value of 0.6 suggests that the difference in age between the groups was not statistically significant. In terms of sex, the data indicate a slightly higher proportion of males in the patient groups than in the control group, although this difference was not statistically significant. The results are compatible with epidemiological data that show that CML patient age differs at the time of analysis, depicting the median age range of 52–64 years.20
According to a different study, CML often manifests in Asians between the ages of 35 and 45, with an annual global incidence rate of 15/1,000,000 and a male-to-female ratio of 1.34. According to several Indian publications, the majority of patients are under 60 years of age.21,22 In their analysis of 856 patients with CML, Berger et al. discovered that women had smaller spleens and higher platelet counts. Additionally, men with CML were less likely to survive and more likely to have additional chromosomal abnormalities. The change was more noticeable among individuals with low and intermediate risk, and it was unrelated to the risk assessment based on the Sokal score.23
Serum PD-L1 protein concentrations were significantly lower in CML patients than in AHC patients. These findings regarding circulating PD-L1 align with the observations of Munari et al. (2021), who reported that circulating PD-L1 levels are influenced by diverse host- and tumor-derived factors and may not consistently correlate with specific gene polymorphisms.24 Collectively, the present results suggest that variation within the PD-L1 gene may contribute to CML pathogenesis through mechanisms distinct from the regulation of circulating protein levels, which exhibit considerable variability.
In the current study, genotypic analysis of the PD-L1 gene revealed distinct differences in the distribution of the CC, CT, and TT genotypes between patients with CML and AHC. The TT genotype was observed in 45% of patients versus 20% of controls, while the T allele was significantly more frequent in patients (60% vs. 25%), suggesting a potential association between the T allele and disease modulation. The CT and TT genotypes demonstrated borderline non-significant p-values (P = 0.08 and P = 0.05, respectively), while the T allele exhibited a statistically significant association with reduced disease risk (P = 0.01, OR = 0.22, CI: 0.06–0.73). These results are partially consistent with those reported previously by Yang M, et al (2024).25 They reported that PD-L1 polymorphisms can influence individual susceptibility to cancer by modulating immune checkpoint signaling. The current findings suggest that the T allele may confer a protective effect by altering the PD-L1/PD-1 interaction, potentially enhancing T cell activity against malignant clones in CML. In contrast, Hashemi M, et al (2019) described a link between PD-L1 overexpression and poor prognosis in hematological malignancies, indicating that PD-L1 regulation may play dual roles, depending on the disease stage and tumor microenvironment.26 Advanced genomic profiling in 2025 identified CD274 (PD-L1) amplification in 19% of CML cases, associated with resistance to second-generation TKIs and poorer survival, highlighting PD-L1 as both a biomarker and a therapeutic target.27
A novel C>T substitution in exon-associated sequences of the PD-L1 gene was identified in both homozygous (TT) and heterozygous (CT) forms. The mutation was confirmed via direct Sanger sequencing and recorded in international genetic databases under accession numbers LC867662, LC867663, and LC867664. Similar mutations have been reported to affect immune surveillance in various malignancies by altering the ligand-receptor binding affinity and expression patterns.28
Importantly, no HBV DNA was detected in the serum of either CML patients or control subjects. This finding is consistent with a study by Kong (2022), which emphasized that HBV reactivation during TKI therapy in CML is rare, particularly in patients without serological signs of prior infection.29 However, this contradicts earlier epidemiological data linking chronic viral infections to leukemia risk through immune modulation and sustained inflammation30. The absence of HBV in the current cohort may be attributed to limited exposure, regional vaccination policies, or the low replication rate of HBV in patients with CML.
The discrepancy in results compared to some previous studies may be attributed to the small sample size, the fact that most patients were receiving treatment, and differences in laboratory equipment or methodology.
The limitations include sample size constraints and lack of functional validation of PD-L1 mutations. Future studies should explore PD-L1 mechanistic role in CML progression and assess the relevance of HBV screening in populations with a higher seroprevalence. These findings underscore PD-L1 genotyping as a potential tool for risk stratification of patients with Iraqi CML.
This study demonstrated a significant reduction in serum PD-L1 levels among Iraqi CML patients compared with healthy controls, suggesting dysregulation of immune checkpoint pathways independent of genetic polymorphisms. The identification of a novel PD-L1 exon mutation (C>T) revealed a potential protective role for the T allele, with carriers exhibiting reduced disease susceptibility. This aligns with PD-L1’s established role in modulating T cell-mediated antitumor responses, where specific variants may enhance immune surveillance against leukemic clones.
7. Data availability
The numerical data generated during this research is available with the authors.
8. Conflict of interest
All authors declare that there was no conflict of interest.
9. Funding
The study utilized the hospital resources only, and no external or industry funding was involved.
10. Ethics Statement
This study adhered to the principles of the Declaration of Helsinki. Before sample collection, verbal and written consent was obtained from all patients. A local ethics commission reviewed and approved the study protocol, consent form, and subject information on September 10, 2024, under project number M240902.
11. Authors’ contribution
All of the authors took equal part in the concept, conduct of the research, data collection, and manuscript preparation.
Authors affiliations:
- Dhuha Oday Hassan, Department of Biology, College of Science, University of Babylon, Hillah, Iraq; Email; dhuhaoday646@gmail.com
- Alaa Hassan Jawad, Department of Biology, College of Science, University of Babylon, Hillah, Iraq; Email; sci.alaa.jawad@uobabylon.edu.iq
- Shakir H. Mohammed Al. Alwany, Department of Biology, College of Science, University of Babylon, Hillah, Iraq; Email; sci.shakir.hammad@uobabylon.edu.iq
ABSTRACT
Background & objective: Patients with chronic myeloid leukemia (CML) remain at a risk of hepatitis B virus (HBV) reactivation during tyrosine kinase inhibitor (TKI) therapy. Programmed death-ligand 1 (PD-L1) polymorphisms may influence immune evasion and disease pathogenesis; however, their interaction with HBV in CML, particularly in Iraqi populations, remains uncharted. We conducted this study to investigate PD-L1 genetic variations, serum PD-L1 levels, and HBV virus infection.
Methodology: This was a case-control study of 120 patients with CML and 100 healthy controls (AHC). Serum PD-L1 and HBV markers were quantified by ELISA Genomic DNA was extracted, and specific regions of the PD-L1 gene and HBV DNA were amplified by PCR. PD-L1 amplicons were sequenced to identify genetic variants.
Results: CML patients exhibited significantly lower serum PD-L1 levels than controls (mean 268.8 ± 89 vs. 470.6 ± 288, P = 0.0032). No HBV DNA was detected in any participant. A novel C>T substitution in PD-L1 was identified, with TT and CT genotypes being more frequent in CML patients than in controls. The T allele showed a borderline significant association with a reduced CML risk (P = 0.01, OR = 0.22, 95% CI: 0.06–0.73).
Conclusion: The PD-L1 T allele may confer protection against CML, while suppression of serum PD-L1 suggests distinct regulatory mechanisms. The absence of HBV highlights regional epidemiological patterns. PD-L1 genotyping can inform risk stratification of patients with Iraqi CML.
Abbreviations: CML: chronic myeloid leukemia, HBV: hepatitis B virus, PD-L1: TKI: tyrosine kinase inhibitor
Keywords: Chronic myeloid leukemia, Programmed Death-Ligand 1, Hepatitis B virus, Immune checkpoint.
Citation: Hassan DO, Jawad AH, Al-Alwany SHM. Immunogenetic study of programmed death-ligand 1 variation and hepatitis B virus in Iraqi patients with chronic myeloid leukemia. Anaesth. pain intensive care 2025;29(7):749-755. DOI: 10.35975/apic.v29i7.2967
Received: May 09, 2025; Revised: October 26, 2025; Accepted: January 01, 2025
1. INTRODUCTION
Chronic myeloid leukemia (CML) is a hematopoietic stem cell (HSC) disorder, like all leukemias, and is a type of cancer that starts in the blood-forming cells of the bone marrow and invades the blood. It is also known as chronic myelogenous leukemia. It is a myeloproliferative neoplasm with an incidence of 1–2 cases per 100,000 adults.1 It is defined by the presence of the Philadelphia chromosome (Ph), which results from reciprocal translocation between chromosomes 9 and 22 [t (9;22] in a single bone marrow cell during cell division. Part of chromosome 9 attaches to chromosome 22, and part of chromosome 22 attaches to chromosome 9. Chromosome 9 was longer than normal, and chromosome 22 was shorter than normal. Abnormal chromosome 22 is known as the “Philadelphia chromosome, which gives rise to a BCR-ABL1 fusion gene.2
CML is classified into three distinct clinical phases: the chronic, accelerated, and blast phases, which are defined by the proportion of immature myeloid blasts in the blood and bone marrow, along with secondary cytogenetic or clinical features.3,4 In the chronic phase, patients typically exhibit <10% blasts and retain partial myeloid differentiation, often presenting with leukocytosis, thrombocytosis, and mild anemia.5,6 Progression to the accelerated phase is marked by 10–19% blasts, ≥20% peripheral basophils, or clonal cytogenetic evolution, reflecting increased genomic instability.7 The blast phase, resembling acute leukemia, is characterized by ≥20% blasts, extramedullary proliferation, or lymphoid blast crisis, indicating a complete loss of differentiation capacity.8
Programmed death ligand 1 (PD-L1), also called (CD274) located on chromosome 9p24.1 and is expressed in non-hematopoietic cells, hematopoietic cells, and antigen-presenting cells (APC).9 An important role of PD-L1 is in various malignancies, where it can attenuate the host immune response to tumor cells.10 PD-L1 expressed by cancer cells binds to PD-1 on the surface of T cells, thereby inhibiting T cell activation and leading to cancer immunity, which allows the tumor to grow without being attacked by the immune system.11
Hepatitis B virus (HBV) infection lasts 45-180 days, most commonly 60-90 days.12 It contains a circular, relaxed DNA (rcDNA) of approximately 3.2 kb in length with a complete minus strand and an incomplete plus strand, which represents the smallest DNA viruses infecting humans. Infected blood or other bodily fluids are involved in the entry of the virus into the body. Contaminated needles or medical equipment, contaminated blood transfusions, unprotected intercourse with an infected individual, or vertical transmission from mother to child during pregnancy or delivery are all possible routes of transmission.13 According to several studies, cytotoxic chemotherapy and immunosuppressive drugs have the potential to hasten the reactivation of chronic HBV infection, putting patients undergoing these treatments at an increased risk for adverse effects.14
Despite established evidence that patients with CML treated with tyrosine kinase inhibitors (TKIs) are at risk of HBV reactivation, standardized HBV screening or monitoring protocols are lacking. Furthermore, while PD-L1 is a critical immune checkpoint implicated in cancer immune evasion, the role of specific PD-L1 gene polymorphism and their potential interaction with HBV infection in modulating CML susceptibility or pathogenesis remains poorly characterized, particularly within the Iraqi population.15,16
This study aimed to address these gaps by investigating PD-L1 genetic variations, PD-L1 serum levels, and HBV infection status as potential risk or protective factors in Iraqi patients with CML
2. METHODOLOGY
This study employed a cross-sectional case-control design. A total of 220 serum specimens were collected, comprising 120 patients diagnosed with CML, and 100 apparently healthy control (AHC) individuals. CML patients’ specimens were obtained from the Iraqi Hematology Center, whereas AHC specimens were collected from general hospitals within the Middle Euphrates region of Iraq. CML patients (aged 20–60 years) met the following criteria: (1) BCR-ABL1+ chronic-phase diagnosis, (2) no active infections at enrollment, (3) no prior allogeneic transplantation, and (4) absence of HIV/HBV/HCV co-infection or concurrent malignancies. AHCs were screened via questionnaire, physical examination, complete blood count (CBC), and C-reactive protein (CRP) testing to exclude immunocompromised states, chronic diseases, or recent infections. All specimens were collected between October 2024 and January 2025.
For serum preparation, approximately 3 mL of venous blood was drawn from each participant into gel-containing serum separators. Samples were centrifuged according to standard protocols to separate the serum, which was then immediately aliquoted and stored at -20°C until subsequent analysis. The serum levels of HBV markers and PD-L1) were quantitatively assessed for both the CML group and the AHC group, using commercially available Enzyme-Linked Immunosorbent Assay (ELISA) kits (BT LAB, China), following the manufacturer's instructions.
2.1. DNA Extraction
Total genomic DNA was isolated from approximately 5 mg of whole blood collected from CML and AHC patients. Samples were immediately transferred to sterile tubes, refrigerated, and homogenized using a hand-held homogenizer in 1.5 mL Eppendorf tubes. Extraction was performed using the G-Spin Total DNA Extraction Kit (iNtRON Biotechnology Co., Korea), and the purified DNA was stored at −20°C. Viral DNA was isolated using a Viral DNA Extraction Kit (iNtRON Biotechnology Co., Korea) under identical storage conditions.
2.2. Primer Design and PCR Amplification
Gene-specific primers were designed for PD-L1 (forward: 5′-TTCCCACTCCCCACTGTTAG-3′; reverse: 5′-GCCCACAGCCACATAAACTT-3′) and HBV (forward: 5′-ACATGGAGAACATCGCATCA-3′; reverse: 5′-AGGACAAACGGGCAACATAC-3′). PCR reactions were conducted in a 25 μL volume comprising 12.5 μL master mix, 5 μL DNA template, 1 μL of each primer, and 5.5 μL nuclease-free water. Amplification was performed using a conventional thermal cycler (Germany) under uniform conditions for both targets: initial denaturation at 95°C for 5 min; 35 cycles of denaturation (95°C for 1 min), annealing (58°C for 45 s), and extension (72°C for 2 min), followed by a final extension at 72°C for 5 min.
Amplicons were resolved using 1.5% agarose gel electrophoresis and visualized under UV transillumination. The PD-L1 PCR products were sequenced using the forward primer via automated Sanger sequencing (Macrogen Inc., Seoul, South Korea). Sequence alignment and analysis were performed using Geneious Bioinformatics software (v2.0) against NCBI reference databases.
2.3. Statistical analysis
To assess the significance of the variables examined in this study, the chi-square test was used. All statistical analyses were conducted using SPSS program version 23. Statistical significance was set at P < 0.05. Hardy-Weinberg equilibrium was used to analyze PD-L1 gene polymorphism.
3. RESULTS
Table 1 compares the demographic and clinical parameters of the 120 CML patients (including 20 newly diagnosed and 100 treated cases) and 100 AHC. Age and sex distributions showed no significant differences (P = 0.06 for age; P = 0.059 for sex), with mean ages of 44 ± 13.3 years (new CML), 48 ± 11.43 years (treated CML), and 46 ± 12.9 years (AHC). However, newly diagnosed CML patients exhibited markedly elevated white blood cell (WBC) counts (173.5 ± 88.13 ×10⁹/L) compared to AHC (6.6 ± 0.75 ×10⁹/L; p<0.001). Platelet counts were also significantly higher in new CML patients (283 ± 138 ×10⁹/L) versus AHC (246.6 ± 56.3 ×10⁹/L; P < 0.05). Hemoglobin (Hb), packed cell volume (PCV), and BCR-ABL ratios did not differ significantly between the CML and AHC groups.
| Table 1: Baseline characteristics of CML patients and healthy controls | |||||
| Parameters | Patients (n = 120) | Controls
(n = 100) |
P-Value | ||
| New diagnosis (n = 20) | Treated
(n = 100) |
||||
| Age (years) | 44 ± 13.3 | 48 ± 11.43 | 46 ± 12.9 | 0.06 | |
| Gender | M: 12 (60) | 52 (52%) | 56 (56%) | 56 (56%) | 0.059 |
| F: 8 (40) | 48 (48%) | 44 (44%) | 44 (44%) | ||
| Gender (M:F) ratio | 1.5:1 | 1.08 | 1.27:1 | ||
| WBC (x109 /L) | 173.5 ± 88.13 | 7.7 ± 5.47 | 6.6 ± 0.75 | < 0.001 | |
| Platelets (x109 /L) | 283 ± 138 | 323 ± 312 | 246.6 ± 56.3 | < 0.05 | |
| Hb (g/dL) | 12.1 ± 1.65 | 12.3 ± 1.65 | 13.1 ± 1.4 | 0.2 | |
| PCV (%) | 36.9 ± 5.21 | 37.5 ± 6.9 | 40.1 ± 4.5 | 0.09 | |
| BCR-ABL ratio | 20.6 ± 6.8 | 6.3 ± 12.6 | 0 | ||
| Data presented as n (%), or mean ± SD: P < 0.05 considered as significant | |||||
Table 2 shows significantly lower serum PD-L1 levels in CML patients (n = 120) than in apparently healthy controls (AHC; n = 100). The mean CTLA-4 concentration was 268.8 ± 89 in CML patients versus 470.6 ± 288 in the AHC group (P = 0.0032).
| Table 2: The serum PD-L1 levels in CML patients and AHC groups | |||
| Groups study | No. of cases | PD-L1 | Groups study |
| Patients | 120 | 268.8 ± 89 | 0.0032 |
| AHC | 100 | 470.6 ± 288 | |
HBV-DNA was absent in all 120 CML patients tested via PCR. All samples were negative (100%), with no positive cases (0%). The P = 0.03 was inconsistent with the non-significant notation (Sign > 0.05), though the data confirm no HBV infection in the cohort. The AHC groups similarly tested negative, supporting the conclusion that HBV was undetectable in both study populations (Figure 1).

Figure 1: non- PCR detection of the HBV gene; in CML patients. M: A 100–1100 bp DNA ladder. After migrating into 2% agarose at 75V and 20 mA for 120 minutes, the PCR-amplified products were stained with ethidium bromide and placed in 15 µl per well
In the current study, the PD-L1 genetic sequences located on chromosome 9 were targeted. The 439 bp amplicon analyzed in this study represents part of the exon region of the PD-L1 molecule (Figure 2).

Figure 2: PCR detection of the PDL-1 gene; in CML patients. M: A 100–1100 bp DNA ladder. After migrating into 2% agarose at 75V and 20 mA for 120 minutes, the PCR-amplified products were stained with ethidium bromide and placed in 15 µl per well.
The NCBI BLASTn engine exhibited approximately 99.9% sequence similarity between the intended target reference sequences and the sample gene sequences. The alignment results of the 493 bp samples were compared to the reference DNA sequences (Figure 3). The mutation was observed at a specific position showing a C > T substitution.

Figure 3: A Novel mutation in PD-L1 gene.
As illustrated in Figure 3, the polymorphism appeared in two distinct zygosity patterns among the sequenced samples: a homozygous (T/T) form and a heterozygous (C/T) form, which was confirmed through sequencing chromatograms. The chromatographic profiles demonstrated a clear peak separation, highlighting the single-nucleotide variation between the samples.
The present results showed that the DNA polymorphism distributions were as follows: CC, CT, and TT were 25%, 30%, and 45%, respectively, in patients with CML and 70%, 10%, and 20%, respectively, in the control group. There were no statistically significant differences (p< 0.05) between the different groups according to PD-L1 genotyping. Statistical analysis of CT indicated a borderline non-significant association (P = 0.08) with an odds ratio (OR) of 0.11 (95% confidence interval [CI]: 0.01–1.32), as shown in Table 4. Statistical analysis of the TT genotype indicated a borderline non-significant association (P = 0.05) with an OR of 0.15 (95% CI: 0.02–1.07), suggesting a potential but not statistically confirmed protective effect.
The C allele was present in 16 patients (40%) and 15 control alleles (75%) and served as the reference allele. The T allele was observed in 24 patients (60%) and 5 control alleles (25%). Statistical analysis showed a borderline significant association between the T allele and decreased disease risk (P = 0.01), with an OR of 0.22 (95% CI: 0.06–0.73) as shown in Table 3.
| Table 3: Genotyping of PD-L1 gene in CML patients and AHC groups | |||||
| Genotype PDL-1 |
Patients
N (%) |
Control
N (%) |
z- statics | P-value | OR (95%)
|
| CC a | 5 (25%) | 7 (70%) | References | ||
| CT | 6 (30%) | 1 (10%) | 1.73 | 0.08 | 0.11 (0.01-1.32) |
| TT | 9 (45%) | 2 (20%) | 1.88 | 0.05* | 0.15 (0.02-1.07) |
| Total number | 20 | 10 | |||
| C allele | 16 (40%) | 15 (75%) | References | ||
| T allele | 24 (60%) | 5 (25%) | 2.47 | 0.01* | 0.22(0.06-0.73) |
| Data presented as n (%); P < 0.05 considered as significant; OR = Odds Ratio | |||||
A novel partial sequence of the PD-L1 gene was identified, covering 439 bp and located on chromosome 9. The samples were collected from blood-derived leukocytes and submitted to the DDBJ/EMBL/GenBank databases under accession numbers LC867662, LC867663, and LC867664.
In this study, the genetic sequence of PD-L1 was analyzed. For the currently investigated 439 bp gene amplicon, BLASTn analysis revealed a high similarity with the reference human PD-L1 gene sequences available in NCBI, including coding and adjacent regulatory regions, according to the human genome annotation (GenBank version: LC867662.1: LC867663.1, and LC867664.1).
4. DISCUSSION
CML is the most common leukemia, in which the BCR-ABL1 fusion gene, which increases tyrosine kinase activity and inhibits apoptotic mechanisms, is responsible for the unchecked clonal proliferation of hematopoietic stem cells that characterizes CML, a myeloproliferative neoplasm. The hallmark of disease pathogenesis is the increase of immature granulocytes in the bone marrow and peripheral blood as a result of this.17–19
In the current study, the average age of the patients with a new diagnosis was 44 years, and the group had an average age of 51 years. However, the P-value of 0.6 suggests that the difference in age between the groups was not statistically significant. In terms of sex, the data indicate a slightly higher proportion of males in the patient groups than in the control group, although this difference was not statistically significant. The results are compatible with epidemiological data that show that CML patient age differs at the time of analysis, depicting the median age range of 52–64 years.20
According to a different study, CML often manifests in Asians between the ages of 35 and 45, with an annual global incidence rate of 15/1,000,000 and a male-to-female ratio of 1.34. According to several Indian publications, the majority of patients are under 60 years of age.21,22 In their analysis of 856 patients with CML, Berger et al. discovered that women had smaller spleens and higher platelet counts. Additionally, men with CML were less likely to survive and more likely to have additional chromosomal abnormalities. The change was more noticeable among individuals with low and intermediate risk, and it was unrelated to the risk assessment based on the Sokal score.23
Serum PD-L1 protein concentrations were significantly lower in CML patients than in AHC patients. These findings regarding circulating PD-L1 align with the observations of Munari et al. (2021), who reported that circulating PD-L1 levels are influenced by diverse host- and tumor-derived factors and may not consistently correlate with specific gene polymorphisms.24 Collectively, the present results suggest that variation within the PD-L1 gene may contribute to CML pathogenesis through mechanisms distinct from the regulation of circulating protein levels, which exhibit considerable variability.
In the current study, genotypic analysis of the PD-L1 gene revealed distinct differences in the distribution of the CC, CT, and TT genotypes between patients with CML and AHC. The TT genotype was observed in 45% of patients versus 20% of controls, while the T allele was significantly more frequent in patients (60% vs. 25%), suggesting a potential association between the T allele and disease modulation. The CT and TT genotypes demonstrated borderline non-significant p-values (P = 0.08 and P = 0.05, respectively), while the T allele exhibited a statistically significant association with reduced disease risk (P = 0.01, OR = 0.22, CI: 0.06–0.73). These results are partially consistent with those reported previously by Yang M, et al (2024).25 They reported that PD-L1 polymorphisms can influence individual susceptibility to cancer by modulating immune checkpoint signaling. The current findings suggest that the T allele may confer a protective effect by altering the PD-L1/PD-1 interaction, potentially enhancing T cell activity against malignant clones in CML. In contrast, Hashemi M, et al (2019) described a link between PD-L1 overexpression and poor prognosis in hematological malignancies, indicating that PD-L1 regulation may play dual roles, depending on the disease stage and tumor microenvironment.26 Advanced genomic profiling in 2025 identified CD274 (PD-L1) amplification in 19% of CML cases, associated with resistance to second-generation TKIs and poorer survival, highlighting PD-L1 as both a biomarker and a therapeutic target.27
A novel C>T substitution in exon-associated sequences of the PD-L1 gene was identified in both homozygous (TT) and heterozygous (CT) forms. The mutation was confirmed via direct Sanger sequencing and recorded in international genetic databases under accession numbers LC867662, LC867663, and LC867664. Similar mutations have been reported to affect immune surveillance in various malignancies by altering the ligand-receptor binding affinity and expression patterns.28
Importantly, no HBV DNA was detected in the serum of either CML patients or control subjects. This finding is consistent with a study by Kong (2022), which emphasized that HBV reactivation during TKI therapy in CML is rare, particularly in patients without serological signs of prior infection.29 However, this contradicts earlier epidemiological data linking chronic viral infections to leukemia risk through immune modulation and sustained inflammation30. The absence of HBV in the current cohort may be attributed to limited exposure, regional vaccination policies, or the low replication rate of HBV in patients with CML.
The discrepancy in results compared to some previous studies may be attributed to the small sample size, the fact that most patients were receiving treatment, and differences in laboratory equipment or methodology.
5. LIMITATIONS
The limitations include sample size constraints and lack of functional validation of PD-L1 mutations. Future studies should explore PD-L1 mechanistic role in CML progression and assess the relevance of HBV screening in populations with a higher seroprevalence. These findings underscore PD-L1 genotyping as a potential tool for risk stratification of patients with Iraqi CML.
6. CONCLUSION
This study demonstrated a significant reduction in serum PD-L1 levels among Iraqi CML patients compared with healthy controls, suggesting dysregulation of immune checkpoint pathways independent of genetic polymorphisms. The identification of a novel PD-L1 exon mutation (C>T) revealed a potential protective role for the T allele, with carriers exhibiting reduced disease susceptibility. This aligns with PD-L1’s established role in modulating T cell-mediated antitumor responses, where specific variants may enhance immune surveillance against leukemic clones.
7. Data availability
The numerical data generated during this research is available with the authors.
8. Conflict of interest
All authors declare that there was no conflict of interest.
9. Funding
The study utilized the hospital resources only, and no external or industry funding was involved.
10. Ethics Statement
This study adhered to the principles of the Declaration of Helsinki. Before sample collection, verbal and written consent was obtained from all patients. A local ethics commission reviewed and approved the study protocol, consent form, and subject information on September 10, 2024, under project number M240902.
11. Authors’ contribution
All of the authors took equal part in the concept, conduct of the research, data collection, and manuscript preparation.
12. REFERENCES
- Jabbour E, Kantarjian H. Chronic myeloid leukemia: 2022 update on diagnosis, therapy, and monitoring. Am J Hematol. 2022;97(9):1236-1256. PMID: 35751859DOI: 1002/ajh.26642
- eininger MW, Shah NP, Altman JK, Berman E, Bhatia R, Bhatnagar B,et al. Chronic myeloid leukemia, version 2.2021, NCCN clinical practice guidelines in oncology. J Natl Compr Cancer Netw. 2020;18(10):1385-1415.PMID: 33022644 DOI: 6004/jnccn.2020.0047
- Rinaldi I, Winston K. Chronic myeloid leukemia, from pathophysiology to treatment-free remission: a narrative literature review. J Blood Med. 2023:261-277. PMCID: PMC10084831 DOI: 2147/JBM.S382090
- \Haznedaroğlu İC, Kuzu I, İlhan O. WHO 2016 Definition of Chronic Myeloid Leukemia and Tyrosine Kinase Inhibitors. Turkish J Haematol Off J Turkish Soc Haematol. 2020;37(1):42-47. PMCID: PMC7057757 DOI: 4274/tjh.galenos.2019.2019.0241
- Senapati J, Jabbour E, Kantarjian H, Short NJ. Pathogenesis and management of accelerated and blast phases of chronic myeloid leukemia. Leukemia. 2023;37(1):5-17.
- Chen WW, Chu TSM, Xu L, et al. Immune related biomarkers for cancer metastasis to the brain. Exp Hematol Oncol. 2022;11(1):105.
- Bonifacio M, Stagno F, Scaffidi L, Krampera M, Di Raimondo F. Management of Chronic Myeloid Leukemia in Advanced Phase. Front Oncol. 2019;9:1132. PMCID: PMC6823861 DOI: 3389/fonc.2019.01132
- Jabbour E, Kantarjian H. Chronic myeloid leukemia: 2025 update on diagnosis, therapy, and monitoring. Am J Hematol. 2024;99(11):2191-2212. PMID: 39093014 DOI: 1002/ajh.27443
- Han Y, Liu D, Li L. PD-1/PD-L1 pathway: current researches in cancer. Am J Cancer Res. 2020;10(3):727.PMCID: PMC7136921
- Lin X, Kang K, Chen P, Zeng Z, Li G, Xiong W, et al. Regulatory mechanisms of PD-1/PD-L1 in cancers. Mol Cancer. 2024;23(1):108. PMCID: PMC11102195 DOI: 1186/s12943-024-02023-w
- Mohajan HK. Prevention of Hepatitis B Virus (HBV) Is Essential to Avoid Chronic Liver Disease. Innov Sci Technol. 2025;4(4):112-121. Available from: https://www.paradigmpress.org/ist/article/view/1658
- Tsukuda S, Watashi K. Hepatitis B virus biology and life cycle. Antiviral Res. 2020;182:104925. DOI: 1016/j.antiviral.2020.104925
- Hoofnagle JH. Reactivation of hepatitis B. Hepatology. 2009;49(S5):S156-S165. DOI: 1002/hep.22945
- Chen Y, Chen H, Huang W, Chen YH, Hsieh TY, Yang SS, et al. Reactivation of hepatitis B virus infection following rituximab treatment in HBsAg‐negative, HBcAb‐positive rheumatoid arthritis patients: a long‐term, real‐world observation. Int J Rheum Dis. 2019;22(6):1145-1151. DOI: 1111/1756-185X.13582
- Sagnelli C, Pisaturo M, Calò F, Martini S, Sagnelli E, Coppola N. Reactivation of hepatitis B virus infection in patients with hemo-lymphoproliferative diseases, and its prevention. World J Gastroenterol. 2019;25(26):3299. PMCID: PMC6639550 DOI: 3748/wjg.v25.i26.3299
- Peiris MN, Li F, Donoghue DJ. BCR: a promiscuous fusion partner in hematopoietic disorders. Oncotarget. 2019;10(28):2738.
- Kuan JW, Su AT, Leong CF, Osato M, Sashida G. Systematic review of normal subjects harbouring BCR-ABL1 fusion gene. Acta Haematol. 2020;143(2):96-111.
- Zhou T, Medeiros LJ, Hu S. Chronic myeloid leukemia: Beyond BCR-ABL1. Curr Hematol Malig Rep. 2018;13:435-445.
- Algahtani FH, Alqahtany FS. Evaluation and characterisation of Chronic myeloid leukemia and various treatments in Saudi Arabia: A retrospective study. J Infect Public Health. 2020;13(2):295-298.
- Mattar M, Bazarbachi A, Abduljalil O, Francis B, Alam A, Blunk V. Epidemiology, Treatment Trends, and Outcomes of Multiple Myeloma in the Middle East and Africa: A Systematic Review. Clin Hematol Int. 2024;6(1):67. PMCID: PMC11086989 DOI: 46989/001c.92555
- Chaulagain P, Poudel A, Aryal S, Sainatham C, Lutfi F. Global Trends of Burden of Chronic Myeloid Leukemia Based on Socio-Demographic Index (SDI): A Comparative Epidemiological Study. Blood. 2024 Nov 5;144:7918..
- Suttorp M, Millot F, Sembill S, Deutsch H, Metzler M. Definition, epidemiology, pathophysiology, and essential criteria for diagnosis of pediatric chronic myeloid leukemia. Cancers (Basel). 2021;13(4):798. PMCID: PMC7917817 DOI: 3390/cancers13040798
- Toloza MJ, Lincango M, Camacho MF, Ledesma MM, Enrico A, Moiraghi B,et al. Immune checkpoints PD1/PD-L1, TIM3/GAL9 and key immune mediators landscape reveal differential expression dynamics on imatinib response in chronic myeloid leukemia. Ann Hematol. 2024;103(12):5249-5260. PMID: 39505795 DOI: 1007/s00277-024-06074-3
- Yang M, Liu Y, Zheng S, et al. Associations of PD-1 and PD-L1 gene polymorphisms with cancer risk: a meta-analysis based on 50 studies. Aging (Albany NY). 2024;16(7):6068.
- Hashemi M, Karami S, Sarabandi S, et al. Association between PD-1 and PD-L1 polymorphisms and the risk of cancer: a meta-analysis of case-control studies. Cancers (Basel). 2019;11(8):1150.
- Zhao F, Zhang X, Tang Y, Yang H, Pan H, Li B, et al. Engineered PD-L1 co-expression in PD-1 knockout and MAGE-C2-targeting TCR-T cells augments the cytotoxic efficacy toward target cancer cells. Sci Rep. 2025;15(1):11894. PMCID: PMC11976951 DOI: 10.1038/s41598-025-92209-z
- Wu Y, Li M, Meng G, Mq T, Ye J. Immune checkpoint‐related gene polymorphisms are associated with acute myeloid leukemia. Cancer Med. 2023;12(18):18588-18596. PMCID: PMC10557852 DOI: 1002/cam4.6468
- Kong JH, Jang JY, Ko TH, Kang SH, Kim Y. High rate of hepatitis B reactivation during tyrosine kinase inhibitor treatment among patients with chronic myeloid leukemia in Korea. Blood Res. 2022;57(4):290-293. doi:10.5045/br.2022.2022099 PMCID: PMC9812728 DOI: 5045/br.2022.2022099
- Chiu CY, Ahmed S, Thomas SK, et al. Hepatitis B Virus Reactivation in Patients Receiving Bruton Tyrosine Kinase Inhibitors. Clin Lymphoma Myeloma Leu k. 2023;23(8):610-615. DOI:10.1016/j.clml.2023.04.006