Hasanain Akram Zainalbden 1 , Emine Arslan 2 , Ihsan Edan Alsaimary 3
Authors affiliations:
Background & objective: Breast cancer (BC) remains a predominant malignancy in women, with rising incidence in younger populations. The tumor's inflammatory microenvironment plays a pivotal role in disease progression. This study evaluates the expression of three key chemokines— Monocyte Chemoattractant Protein-1 (CCL2), Regulated Upon Activation, Normal T Cell Expressed and Secreted (CCL5), and Interferon Gamma-Induced Protein 10 (CXCL10) —in BC patients, aiming to uncover their diagnostic and prognostic significance.
Methodology: BC patients who participated in this study were 100 people who were selected from the Kirkuk Oncology and Hematology Hospital in Kirkuk Governorate (Iraq) from December 2022 to May 2023, and 100 people were considered as a control group after examining them and making sure that they did not have any breast problems or any other health problems. DNA was extracted using the EasyPure Blood Genomic DNA Kit (Lot #X23T7, BioTrans, USA), and gene amplification was performed using conventional PCR with custom-designed primers validated through Primer-BLAST. PCR products were assessed via 1% agarose gel electrophoresis at 110 V for 15 minutes. Internal control genes (β-actin) were used. Duplicate PCR plating and technical replicates were included for quality control and reproducibility.
Results: Clear amplification of the three chemokine genes was observed: CCL2 (490 bp), CCL5 (481 bp), and CXCL10 (310 bp). Elevated expression was noted in late-stage BC patients. DNA quality was verified using NanoDrop OneC (Thermo Scientific, v1.6.3), and all samples passed quality assurance thresholds (A260/280 ≥ 1.8).
Conclusion: The study highlights the presence of CCL2, CCL5, and CXCL10 in BC and highlights their roles as biomarkers in the BC investigation and therapeutic targets. These results contribute to a better understanding of the molecular mechanisms that form the basis of breast cancer and highlight the value of chemokine profiling in disease management, prognosis, and treatment planning.
Abbreviations: BC: Breast cancer, CCL2: C-C Motif Chemokine Ligand 2, CCL5: C-C Motif Chemokine Ligand 2, CCR: chemokine receptor, CXCL10: C-X-C motif chemokine ligand 10, MCP-1: Monocyte Chemoattractant Protein-1, PCR: polymerase chain reaction
Keywords: Biomarkers; Breast Cancer; Chemokines; Tumor Microenvironment; PCR; Polymerase Chain Reaction
Citation: Zainalbden HA, Arslan E, Alsaimary IE. Expression analysis of CCL2, CCL5, and CXCL10 in breast cancer: an insight into the inflammatory microenvironment. Anaesth. pain intensive care 2025;29(6):513-519. DOI: 10.35975/apic.v29i6.2901
Received: May 09, 2024; Revised: October 26, 2024; Accepted: January 01, 2025
Breast Cancer (BC) is the leading cause of cancer death and the most common cancer among women, with a lifetime diagnosis rate of 9%.1 It constitutes one-third of all reported malignancies in women, making it the most common type of cancer in this population, and the disease is increasingly affecting younger Age categories.2 BC has become one of the greatest threats to the health of Iraqi women over the past two decades, with increasing incidence rates. Chemokines and their receptors are essential in guiding the trafficking of both bone marrow-derived and resident cells to areas of inflammation.3 In addition to their functions in promoting leukocyte recruitment, chemokines also play a role in immune regulation and T cell polarization, the induction of processes such as cell division, angiogenesis, tumor metastasis, respiratory burst, apoptosis, wound healing, and proteins from the extracellular matrix and the secretion of cytokines is crucial in various biological contexts.4 This study investigates the molecular expression of CCL2, CCL5, and CXCL10 in BC patients using conventional PCR, aiming to elucidate their roles in disease pathogenesis and evaluate their utility as diagnostic or prognostic markers.
2.4. DNA Quantification and Quality Check
Extracted DNA was assessed using the NanoDrop OneC Spectrophotometer (Thermo Scientific®, Software Version 1.6.3). DNA purity was determined by A260/A280 ratio, and samples with values between 1.8–2.0 were deemed acceptable. DNA was stored at -20°C until further analysis.
Primer sequences are detailed in Table 3.
2.9. DNA Amplification
PCR (Conventional PCR) was used for the amplification of patients' DNA samples to obtain the targeted genes by using their specific primers as in Table 2, according to the manufacturer's instructions of the DNA extraction kit (EasyPure® Blood Genomic DNA Kit), where used in our study.
3.1. DNA Extraction and Quality Control
High-quality genomic DNA was extracted from the whole blood of the control group and BC patients using the EasyPure® Blood Genomic DNA Kit. Purity ratios (A260/A280) ranged from 1.83 to 1.95, which yields DNA of very high quality and quantity. Each sample was extracted individually and subsequently analyzed by gel electrophoresis to confirm the results, without signs of degradation or contamination, as illustrated in Figure 1.
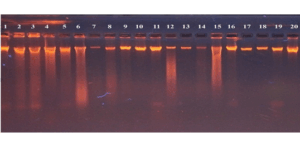
Figure 1: Agrose gel electrophoresis stained with ethidium bromide dye showed clear
bands that represent DNA molecules extracted from whole blood.
3.2. PCR Amplification
DNA extracted from whole blood samples was amplified using conventional PCR, and the results of the PCR products were confirmed by 1% agarose gel electrophoresis (at 110 V for 15 minutes). In this analysis, the DNA bands that appeared on the gel indicated successful binding of the extracted DNA template to the specific primers targeting each of the selected chemokines studied (CCL2, CCL5, and CXCL10). Bands with the expected molecular size of 490 bp were observed, which is specific for the CCL2 primer, 481 bp for the CCL5 primer, and 310 bp for the CXCL10 primer.
β-actin bands were consistently present across all samples, validating internal control integrity. No bands appeared in the no-template controls, confirming the absence of contamination. As shown in Figures 2, 3, and 4.
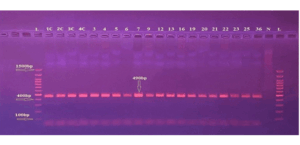
Figure 2: Gel electrophoresis for PCR product of CCL2 primer shows 490 bp Primer: DNA
ladder (100-1500 bp). Lane N represents the negative control, and Lanes (IC-36) represent
a positive result
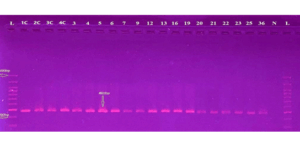
Figure 3: Gel electrophoresis for PCR product of CCL5 primer shows 481 bp Primer L: DNA
ladder (100-1500 bp). Lane N represents the negative control, and Lanes (IC-36) represent
a positive result
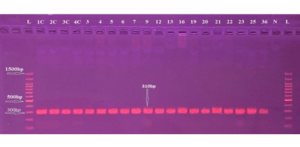
Figure 4: Gel electrophoresis for PCR product of CXCL10 primer shows 310 bp Primer L: DNA Ladder (100-1500 bp), lane N represents the negative control, and Lanes (IC-36) represent the positive results
3.3. Reproducibility and Replication
Duplicate plating of each DNA sample in PCR assays confirmed reproducibility. Technical replicates yielded consistent amplification results across multiple runs. Select samples were re-extracted and re-amplified to validate the stability of the chemokine expression signals.
This study provides compelling evidence for the elevated expression of three inflammatory chemokines—Monocyte Chemoattractant Protein-1 (CCL2), Regulated Upon Activation, Normal T Cell Expressed and Secreted (CCL5), and Interferon Gamma-Induced Protein 10 (CXCL10)—in breast cancer (BC) patients compared to healthy controls. Notably, expression levels were significantly higher in patients with advanced-stage disease, suggesting a correlation between chemokine expression and tumor progression.
CCL2, also known as MCP-1, is a pivotal chemokine responsible for recruiting monocytes to sites of inflammation. In cancer, it has been implicated in supporting tumor-associated macrophages (TAMs), enhancing metastasis, and promoting an immunosuppressive environment.5,6 Our findings align with Liu et al., who reported elevated CCL2 expression in basal-like BC, contributing to tumor growth via fibroblast activation and immune modulation.6
CCL5, commonly referred to as RANTES, also showed increased amplification in our study. CCL5 is produced by T lymphocytes and dendritic cells and interacts with CCR1, CCR3, and CCR5 receptors, thereby facilitating immune cell recruitment and cytokine secretion.7,8 Research has shown that CCL5 promotes epithelial-mesenchymal transition (EMT), a key mechanism in cancer metastasis. In BC patients, high plasma levels of CCL5 were associated with increased expression of CX3CL1 and activation of epidermal growth factor pathways.9 These insights are consistent with our data, further reinforcing CCL5’s role in aggressive tumor phenotypes.
CXCL10 (IP-10) is induced by interferon-gamma and primarily interacts with the CXCR3 receptor. It recruits CD8+ cytotoxic T cells and Th1 cells, potentially enhancing anti-tumor immunity. However, its role in BC remains paradoxical. While it can enhance cytotoxic T cell infiltration, chronic CXCL10 expression may also create a pro-inflammatory milieu that fosters tumor growth.10,11 In this study, the observed upregulation of CXCL10 suggests its dual role—immune recruitment and potential immunosuppression in a chronically inflamed microenvironment.
A retrospective strength of this study lies in the utilization of archived blood samples processed under standardized protocols (STP), quality-verified DNA, and reproducibility ensured by technical duplicates and blinded PCR runs. The strong concordance of molecular data across biological replicates enhances the robustness of these findings.
Furthermore, the integration of quality assurance (QA) and quality control (QC) practices—including lot-controlled reagents, calibrated instruments, and internal control validation—provides confidence in the molecular results. The absence of amplification in negative controls and the consistent presence of β-actin bands affirm the specificity and integrity of the experimental workflow.
Comparing our findings with existing literature also underscores the potential of these chemokines as multi-biomarker panels for BC diagnosis and prognosis. No prior studies, to our knowledge, have simultaneously evaluated CCL2, CCL5, and CXCL10 using conventional PCR from blood-derived DNA in a Middle Eastern cohort, thereby offering novel geographical and ethnic insight.
In summary, the upregulated expression of these chemokines emphasizes their potential as non-invasive biomarkers and therapeutic targets. Future work should involve quantitative PCR (qPCR) for expression quantification and longitudinal studies to monitor changes in chemokine levels during treatment.
This study underscores the molecular significance of the chemokines Monocyte Chemoattractant Protein-1 (CCL2), Regulated Upon Activation, Normal T Cell Expressed and Secreted (CCL5), and Interferon Gamma-Induced Protein 10 (CXCL10) in the inflammatory microenvironment of breast cancer (BC). Elevated expression of these genes in BC patients, particularly those with advanced-stage disease, highlights their involvement in tumor progression, immune modulation, and potential metastasis.
Through rigorous quality assurance, validated primer design, internal control application, and technical replication, the study provides a robust molecular basis for chemokine profiling. These findings contribute to growing evidence that chemokines are not only key mediators of inflammation but also viable biomarkers for diagnosis, prognosis, and possibly targets for therapeutic intervention in breast cancer.
6. Future investigations
Future investigations should employ quantitative PCR (qPCR) to evaluate relative gene expression levels, explore serum protein correlations via ELISA, and assess the predictive value of these chemokines in response to therapy.
7. Data availability
The numerical data generated during this research are available from the authors.
8. Conflict of interest
All authors declare that there was no conflict of interest.
9. Funding
The study utilized the hospital resources only, and no external or industry funding was involved.
10. Acknowledgements
The authors express their sincere gratitude to the staff of the Department of Biology, Faculty of Science, Selcuk University, Konya, Turkey, and the Department of Microbiology, College of Medicine, University of Basra, Basra, Iraq.
11. Authors’ contribution
HAZ: Concept, study design, data analysis, manuscript writing, and final approval
EMA: First advisor, literature review, manuscript editing
IHA: Second advisor, data interpretation, supervision, manuscript revision
Authors affiliations:
- Hasanain Akram Zainalbden, Department of Biology, Faculty of Science, Selcuk University, Konya, Turkey / Tuz General Hospital, Saladin Health Directorate, Ministry of Health, Iraq; Email: hasanainakram1981@gmail.com
- Emine Arslan, Department of Biology, Faculty of Science, Selcuk University, Konya, Turkey; Email: earslan@selcuk.edu.tr
- Ihsan Edan Alsaimary, Department of Microbiology, College of Medicine, University of Basra, Basra, Iraq; Email: ihsan.alsaimary@uobasrah.edu.iq
ABSTRACT
Background & objective: Breast cancer (BC) remains a predominant malignancy in women, with rising incidence in younger populations. The tumor's inflammatory microenvironment plays a pivotal role in disease progression. This study evaluates the expression of three key chemokines— Monocyte Chemoattractant Protein-1 (CCL2), Regulated Upon Activation, Normal T Cell Expressed and Secreted (CCL5), and Interferon Gamma-Induced Protein 10 (CXCL10) —in BC patients, aiming to uncover their diagnostic and prognostic significance.
Methodology: BC patients who participated in this study were 100 people who were selected from the Kirkuk Oncology and Hematology Hospital in Kirkuk Governorate (Iraq) from December 2022 to May 2023, and 100 people were considered as a control group after examining them and making sure that they did not have any breast problems or any other health problems. DNA was extracted using the EasyPure Blood Genomic DNA Kit (Lot #X23T7, BioTrans, USA), and gene amplification was performed using conventional PCR with custom-designed primers validated through Primer-BLAST. PCR products were assessed via 1% agarose gel electrophoresis at 110 V for 15 minutes. Internal control genes (β-actin) were used. Duplicate PCR plating and technical replicates were included for quality control and reproducibility.
Results: Clear amplification of the three chemokine genes was observed: CCL2 (490 bp), CCL5 (481 bp), and CXCL10 (310 bp). Elevated expression was noted in late-stage BC patients. DNA quality was verified using NanoDrop OneC (Thermo Scientific, v1.6.3), and all samples passed quality assurance thresholds (A260/280 ≥ 1.8).
Conclusion: The study highlights the presence of CCL2, CCL5, and CXCL10 in BC and highlights their roles as biomarkers in the BC investigation and therapeutic targets. These results contribute to a better understanding of the molecular mechanisms that form the basis of breast cancer and highlight the value of chemokine profiling in disease management, prognosis, and treatment planning.
Abbreviations: BC: Breast cancer, CCL2: C-C Motif Chemokine Ligand 2, CCL5: C-C Motif Chemokine Ligand 2, CCR: chemokine receptor, CXCL10: C-X-C motif chemokine ligand 10, MCP-1: Monocyte Chemoattractant Protein-1, PCR: polymerase chain reaction
Keywords: Biomarkers; Breast Cancer; Chemokines; Tumor Microenvironment; PCR; Polymerase Chain Reaction
Citation: Zainalbden HA, Arslan E, Alsaimary IE. Expression analysis of CCL2, CCL5, and CXCL10 in breast cancer: an insight into the inflammatory microenvironment. Anaesth. pain intensive care 2025;29(6):513-519. DOI: 10.35975/apic.v29i6.2901
Received: May 09, 2024; Revised: October 26, 2024; Accepted: January 01, 2025
1. INTRODUCTION
Breast Cancer (BC) is the leading cause of cancer death and the most common cancer among women, with a lifetime diagnosis rate of 9%.1 It constitutes one-third of all reported malignancies in women, making it the most common type of cancer in this population, and the disease is increasingly affecting younger Age categories.2 BC has become one of the greatest threats to the health of Iraqi women over the past two decades, with increasing incidence rates. Chemokines and their receptors are essential in guiding the trafficking of both bone marrow-derived and resident cells to areas of inflammation.3 In addition to their functions in promoting leukocyte recruitment, chemokines also play a role in immune regulation and T cell polarization, the induction of processes such as cell division, angiogenesis, tumor metastasis, respiratory burst, apoptosis, wound healing, and proteins from the extracellular matrix and the secretion of cytokines is crucial in various biological contexts.4 This study investigates the molecular expression of CCL2, CCL5, and CXCL10 in BC patients using conventional PCR, aiming to elucidate their roles in disease pathogenesis and evaluate their utility as diagnostic or prognostic markers.
2. METHODOLOGY
2.1. Study Population and Sample Collection
In this study, BC cases (case/control study) were collected in Kirkuk province, especially those who applied to Kirkuk Oncology and Hematology Hospital (Kirkuk, Iraq) between December 2022 and May 2023. All medical information of the patients, including name, age, residence, and marital status, was recorded on the questionnaire sheet, including women with early-stage, second-stage, and last-stage BC after receiving all treatments. All participants gave informed consent. All information was collected from the patients after the oncologist made the correct diagnosis and confirmed it with other findings (Cystoscopy, biopsy, ultrasound, and CT scan).2.2. STP for sample collection
Whole blood (5 mL) was collected in sterile EDTA-coated vacutainer tubes. Samples were immediately stored at 4°C and processed within 6 hours. For longer-term storage, samples were frozen at -80°C until DNA extraction.2.3. DNA Extraction
Genomic DNA was extracted from 5 mL frozen whole blood using the EasyPure® Blood Genomic DNA Kit (BioTrans®, USA; Lot No X23T7) simply and conveniently. To liberate DNA, whole blood is treated with binding/lysis buffer. and performed according to the manufacturer's procedural directive (Table 1). Each extraction included an internal reference control (IRC) to monitor the procedure's integrity. Negative controls (without template DNA) were incorporated in each batch to rule out contamination.| Table 1: DNA Extraction Components Kit | |
| Components | Quantity |
| Binding Buffer 3 (BB3) | 30 mL |
| Clean Buffer 3 (CB3) | 6 mL |
| Wash Buffer 3 (WB3) | 12 mL |
| Elution Buffer (EB) | 25 mL |
| RNase A (20 mg/mL) | 500 μl (EE121-01) |
| 0 (EE121-11) | |
| Proteinase K (20 mg/mL) | 1 mL |
| Genomic Spin Columns with Collection Tubes | 50 each |
2.4. DNA Quantification and Quality Check
Extracted DNA was assessed using the NanoDrop OneC Spectrophotometer (Thermo Scientific®, Software Version 1.6.3). DNA purity was determined by A260/A280 ratio, and samples with values between 1.8–2.0 were deemed acceptable. DNA was stored at -20°C until further analysis.
2.5. Designing of Primer
Primers for CCL2, CCL5, and CXCL10 were designed using NCBI Primer-BLAST. Primer pairs were selected to ensure high specificity and minimal dimer formation. The primer design criteria included target fragment size: CCL2 (490 bp), CCL5 (481 bp), CXCL10 (310 bp), melting temperature (Tm) between 58–62°C, and GC percentage between 45–55%.| Table 2: PCR master mix volume | |
| Components | Volume |
| 10 µM Forward Primer | 0.5 µL |
| 10 µM Reverse Primer | 0.5 µL |
| Template DNA | 1.5 µL |
| OneTaq Quick-Load 2X Master Mix with Standard Buffer |
12.5 µL |
| Nuclease-free water | 10 µL |
| Total | 25 µL |
Primer sequences are detailed in Table 3.
| Table 3: Primer sequences and product size | |||
| References | Amplicon size | Primer sequences (5'-'3) | Target gee |
| Gene bank accession number: LC727557 | 490bp | F: AGAATCACCAGCAGCAAGTGTCC R: TCCTGAACCCACTTCTGCTTGG |
CCL2 |
| Gene bank accession number: LC727558 | 481bp | F: ACACTTGACATTGTGCTGGAC R: AGTGGCAACTGATGCTTCCC |
CCL5 |
| Gene bank accession number: LC727559 | 310bp | F: CCAATTTTGTCCACGTGTTG R: TTCTTGATGGCCTTCGATTC |
CXCL10 |
2.6. PCR Amplification
A conventional PCR reaction was performed in a 25 µL volume using specific components and a standard thermal protocol, including denaturation, annealing, and extension steps, with negative controls and the β-actin gene included as an internal control.2.7. Gel Electrophoresis (GE)
PCR products were analyzed on 1% agarose gels prepared with LE Agarose (Bio-Rad®, Lot #LR456789) in 1x TBE buffer. Gels were stained with GelRed™ and visualized using a Gel Doc XR+ Imaging System (Bio-Rad®, Software Version 5.2.1). A 100 bp DNA ladder (Thermo Scientific®) was used for sizing.2.8. Quality Assurance and Reproducibility
All experiments were performed under sterile conditions in a Class II biosafety cabinet with duplicate samples to ensure consistency, random replication, and blind analysis to minimize bias, with instrument calibration regularly documented.| Table 4: Conditions of working | ||||||
| Gene | Temperature (°C) / Time | Cycle No. | ||||
| Initial denaturation | Cycling temperature | Final extension | ||||
| Denaturation | Annealing | Extension | ||||
| CCL2 | 94ºc / 30sec | 94ºc / 30 sec | 45-68ºc / 15-60 sec | 68ºc/1 min | 68ºc / 5 min | 30 |
| CCL5 | 94ºc / 30sec | 94ºc / 30 sec | 45-68ºc / 15-60 sec | 68ºc/1 min | 68ºc / 5 min | 30 |
| CXCL10 | 94ºc / 30sec | 94ºc / 30 sec | 45-68ºc / 15-60 sec | 68ºc/1 min | 68ºc / 5 min | 30 |
2.9. DNA Amplification
PCR (Conventional PCR) was used for the amplification of patients' DNA samples to obtain the targeted genes by using their specific primers as in Table 2, according to the manufacturer's instructions of the DNA extraction kit (EasyPure® Blood Genomic DNA Kit), where used in our study.
3. RESULTS
3.1. DNA Extraction and Quality Control
High-quality genomic DNA was extracted from the whole blood of the control group and BC patients using the EasyPure® Blood Genomic DNA Kit. Purity ratios (A260/A280) ranged from 1.83 to 1.95, which yields DNA of very high quality and quantity. Each sample was extracted individually and subsequently analyzed by gel electrophoresis to confirm the results, without signs of degradation or contamination, as illustrated in Figure 1.

Figure 1: Agrose gel electrophoresis stained with ethidium bromide dye showed clear
bands that represent DNA molecules extracted from whole blood.
3.2. PCR Amplification
DNA extracted from whole blood samples was amplified using conventional PCR, and the results of the PCR products were confirmed by 1% agarose gel electrophoresis (at 110 V for 15 minutes). In this analysis, the DNA bands that appeared on the gel indicated successful binding of the extracted DNA template to the specific primers targeting each of the selected chemokines studied (CCL2, CCL5, and CXCL10). Bands with the expected molecular size of 490 bp were observed, which is specific for the CCL2 primer, 481 bp for the CCL5 primer, and 310 bp for the CXCL10 primer.
β-actin bands were consistently present across all samples, validating internal control integrity. No bands appeared in the no-template controls, confirming the absence of contamination. As shown in Figures 2, 3, and 4.

Figure 2: Gel electrophoresis for PCR product of CCL2 primer shows 490 bp Primer: DNA
ladder (100-1500 bp). Lane N represents the negative control, and Lanes (IC-36) represent
a positive result

Figure 3: Gel electrophoresis for PCR product of CCL5 primer shows 481 bp Primer L: DNA
ladder (100-1500 bp). Lane N represents the negative control, and Lanes (IC-36) represent
a positive result

Figure 4: Gel electrophoresis for PCR product of CXCL10 primer shows 310 bp Primer L: DNA Ladder (100-1500 bp), lane N represents the negative control, and Lanes (IC-36) represent the positive results
3.3. Reproducibility and Replication
Duplicate plating of each DNA sample in PCR assays confirmed reproducibility. Technical replicates yielded consistent amplification results across multiple runs. Select samples were re-extracted and re-amplified to validate the stability of the chemokine expression signals.
4. DISCUSSION
This study provides compelling evidence for the elevated expression of three inflammatory chemokines—Monocyte Chemoattractant Protein-1 (CCL2), Regulated Upon Activation, Normal T Cell Expressed and Secreted (CCL5), and Interferon Gamma-Induced Protein 10 (CXCL10)—in breast cancer (BC) patients compared to healthy controls. Notably, expression levels were significantly higher in patients with advanced-stage disease, suggesting a correlation between chemokine expression and tumor progression.
CCL2, also known as MCP-1, is a pivotal chemokine responsible for recruiting monocytes to sites of inflammation. In cancer, it has been implicated in supporting tumor-associated macrophages (TAMs), enhancing metastasis, and promoting an immunosuppressive environment.5,6 Our findings align with Liu et al., who reported elevated CCL2 expression in basal-like BC, contributing to tumor growth via fibroblast activation and immune modulation.6
CCL5, commonly referred to as RANTES, also showed increased amplification in our study. CCL5 is produced by T lymphocytes and dendritic cells and interacts with CCR1, CCR3, and CCR5 receptors, thereby facilitating immune cell recruitment and cytokine secretion.7,8 Research has shown that CCL5 promotes epithelial-mesenchymal transition (EMT), a key mechanism in cancer metastasis. In BC patients, high plasma levels of CCL5 were associated with increased expression of CX3CL1 and activation of epidermal growth factor pathways.9 These insights are consistent with our data, further reinforcing CCL5’s role in aggressive tumor phenotypes.
CXCL10 (IP-10) is induced by interferon-gamma and primarily interacts with the CXCR3 receptor. It recruits CD8+ cytotoxic T cells and Th1 cells, potentially enhancing anti-tumor immunity. However, its role in BC remains paradoxical. While it can enhance cytotoxic T cell infiltration, chronic CXCL10 expression may also create a pro-inflammatory milieu that fosters tumor growth.10,11 In this study, the observed upregulation of CXCL10 suggests its dual role—immune recruitment and potential immunosuppression in a chronically inflamed microenvironment.
A retrospective strength of this study lies in the utilization of archived blood samples processed under standardized protocols (STP), quality-verified DNA, and reproducibility ensured by technical duplicates and blinded PCR runs. The strong concordance of molecular data across biological replicates enhances the robustness of these findings.
Furthermore, the integration of quality assurance (QA) and quality control (QC) practices—including lot-controlled reagents, calibrated instruments, and internal control validation—provides confidence in the molecular results. The absence of amplification in negative controls and the consistent presence of β-actin bands affirm the specificity and integrity of the experimental workflow.
Comparing our findings with existing literature also underscores the potential of these chemokines as multi-biomarker panels for BC diagnosis and prognosis. No prior studies, to our knowledge, have simultaneously evaluated CCL2, CCL5, and CXCL10 using conventional PCR from blood-derived DNA in a Middle Eastern cohort, thereby offering novel geographical and ethnic insight.
In summary, the upregulated expression of these chemokines emphasizes their potential as non-invasive biomarkers and therapeutic targets. Future work should involve quantitative PCR (qPCR) for expression quantification and longitudinal studies to monitor changes in chemokine levels during treatment.
5. CONCLUSION
This study underscores the molecular significance of the chemokines Monocyte Chemoattractant Protein-1 (CCL2), Regulated Upon Activation, Normal T Cell Expressed and Secreted (CCL5), and Interferon Gamma-Induced Protein 10 (CXCL10) in the inflammatory microenvironment of breast cancer (BC). Elevated expression of these genes in BC patients, particularly those with advanced-stage disease, highlights their involvement in tumor progression, immune modulation, and potential metastasis.
Through rigorous quality assurance, validated primer design, internal control application, and technical replication, the study provides a robust molecular basis for chemokine profiling. These findings contribute to growing evidence that chemokines are not only key mediators of inflammation but also viable biomarkers for diagnosis, prognosis, and possibly targets for therapeutic intervention in breast cancer.
6. Future investigations
Future investigations should employ quantitative PCR (qPCR) to evaluate relative gene expression levels, explore serum protein correlations via ELISA, and assess the predictive value of these chemokines in response to therapy.
7. Data availability
The numerical data generated during this research are available from the authors.
8. Conflict of interest
All authors declare that there was no conflict of interest.
9. Funding
The study utilized the hospital resources only, and no external or industry funding was involved.
10. Acknowledgements
The authors express their sincere gratitude to the staff of the Department of Biology, Faculty of Science, Selcuk University, Konya, Turkey, and the Department of Microbiology, College of Medicine, University of Basra, Basra, Iraq.
11. Authors’ contribution
HAZ: Concept, study design, data analysis, manuscript writing, and final approval
EMA: First advisor, literature review, manuscript editing
IHA: Second advisor, data interpretation, supervision, manuscript revision
12. REFERENCES
- Łukasiewicz S, Czeczelewski M, Forma A, Baj J, Sitarz R, Stanisławek A. Breast cancer— epidemiology, risk factors, classification, prognostic markers, and current treatment strategies— an updated review. Cancers. 2021;13(17):4287. [PubMed] DOI: 3390/cancers13174287
- Zhu J, Charkhchi P, Adekunte S, Akbari MR. What is known about breast cancer in young women? Cancers. 2023;15(6):191-7. [PubMed] DOI: 3390/cancers15061917
- K Interleukin-33 Levels are Up-Regulated in Women with Breast Cancer and Shown to be Associated with the Triple-Negative Type. Iraq J Sci. 2024;65(9):5005-1 DOI: 24996/ijs.2024.65.9.17
- Mir MA, Rashid M, Jan N. Cytokines and Chemokines in Tumor Growth and Progression. In: Cytokine and Chemokine Networks in Cancer. Singapore: Springer Nature Singapore; 2023. p. 33-77.
- Yoshimura T, Li C, Wang Y, Matsukawa A. The chemokine monocyte chemoattractant protein1/CCL2 is a promoter of breast cancer metastasis. Cell Mol Immunol. 2023;20(7):714738. [PubMed] DOI: 1038/s41423-023-01013-0
- Liu H, Yang Z, Lu W, Chen Z, Chen L, Han S, Cai Y. Chemokines and chemokine receptors: A new strategy for breast cancer therapy. Cancer Med. 2020;9(11):3786-3799. [PubMed] DOI: 1002/cam4.3014
- Fırat AA. Chemokines effective on platelet functions. In: Chemokines updates. IntechOpen; 2022. Available from: https://www.intechopen.com/chapters/84395
- Berghi NO, Dumitru M, Vrinceanu D, Ciuluvica RC, Simioniuc-Petrescu A, Caragheorgheopol R, et al. Relationship between chemokines and T lymphocytes in the context of respiratory allergies. Exp Ther Med. 2020;20(3):2352-2360. [PubMed] DOI: 3892/etm.2020.8961
- Li H, Wu M, Zhao X. Role of chemokine systems in cancer and inflammatory diseases. MedComm. 2022;3(2):e147. [PubMed] DOI: 1002/mco2.147
- Rahmat-Zaie R, Amini J, Haddadi M, Beyer C, Sanadgol N, Zendedel A. TNF- α/STAT1/CXCL10 mutual inflammatory axis that contributes to the pathogenesis of experimental models of multiple sclerosis: A promising signaling pathway for targeted therapies. Cytokine. 2023;168:156235. [PubMed] DOI: 1016/j.cyto.2023.156235
- Xiao W, Huang H, Zheng P, Liu Y, Chen Y, Chen J, et al. The CXCL10/CXCR3 pathway contributes to the synergy of thermal ablation and PD-1 blockade therapy against tumors. Cancers. 2023;15(5):1427. [PubMed] DOI: 3390/cancers15051427